Computer Aided Drug Design (CADD) has become increasingly crucial for PROTAC design as the accuracy of theoretical models has improved. The most promising PROTAC molecules for specific targets and E3 ligases can be screened in a relatively short amount of time with the aid of CADD, saving experimentalists a lot of testing time and capital costs. The Viva CADD platform uses a new strategy for PROTAC design that combines techniques reported in the literature on CADD design of PROTAC molecules with Viva's enhanced molecular dynamics sampling technology to further improve the accuracy of computational results. This improvement will help design optimized PROTAC molecules, thus increasing the success rate of PROTAC design.
CADD-Based PROTAC Design Consists of the Following Main Steps:
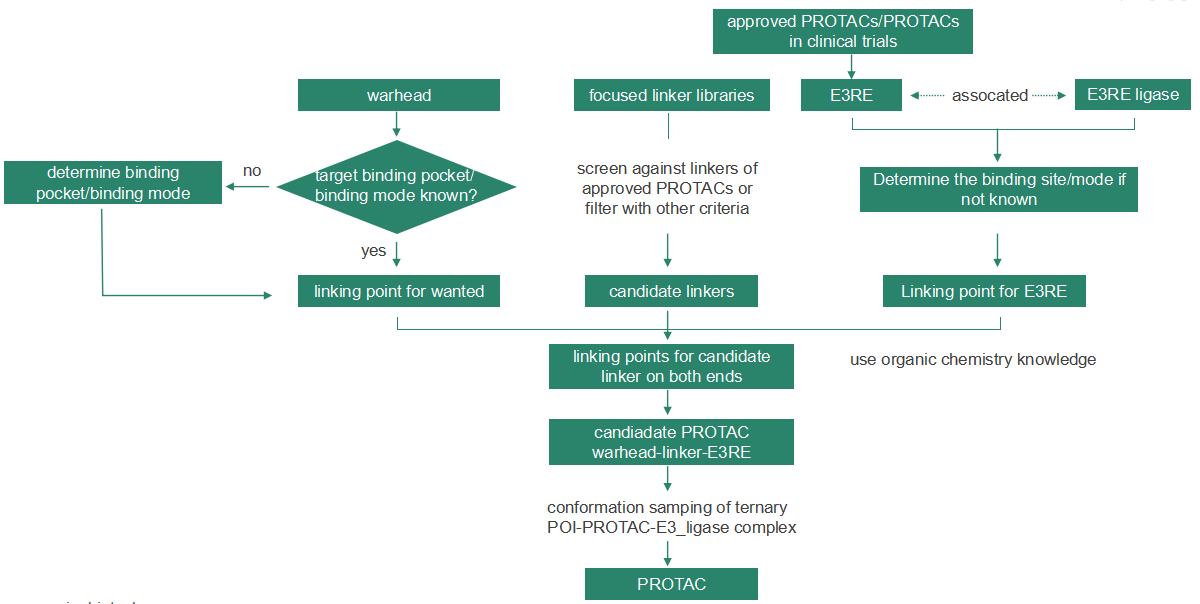
First, the warhead and E3 ligand can be obtained by structure-based ligand design, and then the binding mode of the target protein and warhead can be analyzed to determine the possible binding modes of the linker and warhead. (Usually, the target protein structure is known, and for E3 ligase, compounds that have been reported in the literature can serve as the starting structures.)
The site and mode of binding of the linker to E3 ligand is also available by the same means. A number of representative linker compounds from each of the common linker types reported in literature can be chosen, and the linkers that could generate stable warhead-linker-ligand combinations (i.e., PROTAC molecules) and further lead to stable target-PROTAC-ligase can be selected. The validity of the resulting PROTAC molecules can be tested through further experiments.
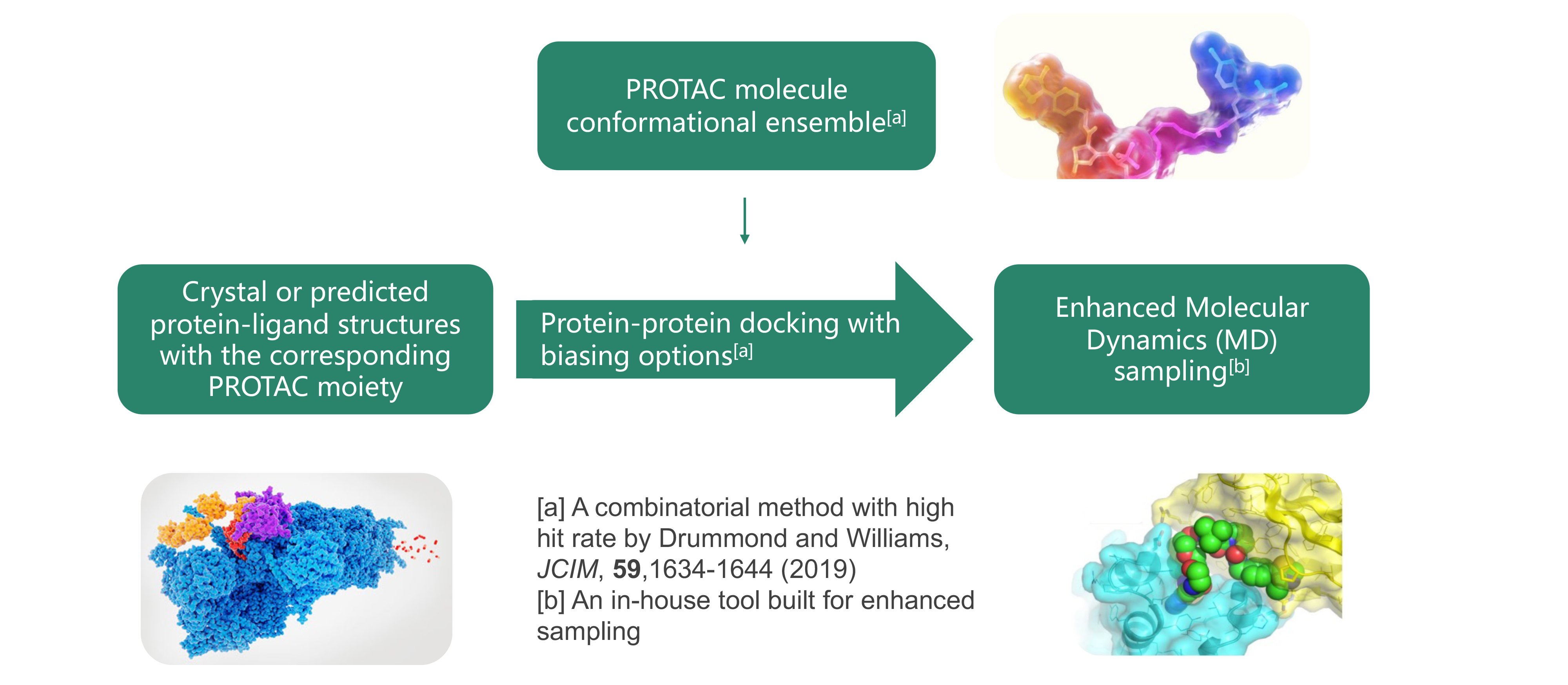
For the conformational analysis of the target-PROTAC-ligase ternary complex, different strategies can be used, such as studying the target-PROTAC first and then the target-PROTAC-ligase, or studying the target-PROTAC-ligase directly.
In Silico PROTAC Design Process: